A major blow to the ’plasticity-first’ hypothesis?
 Skeletal morphology is a classic candidate for ‘plasticity-led’ evolution. Our new paper in eLife tests the idea that such ‘plasticity-led’ evolution has occurred in the adaptive radiation of Anolis lizards on the Greater Antilles.
Skeletal morphology is a classic candidate for ‘plasticity-led’ evolution. Our new paper in eLife tests the idea that such ‘plasticity-led’ evolution has occurred in the adaptive radiation of Anolis lizards on the Greater Antilles.
In her study, Nathalie quantified the morphological divergence between ecomorphs on the four Greater Antillean islands. She then compared this to the morphological changes induced by microhabitats that promote climbing or running. It turns out that divergences between ecomorphs have occurred in parallel on the different islands. However, the divergence between ecomorphs was not aligned with plastic responses. In addition, plasticity differed between species, making it too transient to exercise a persistent effect on adaptive diversification of the locomotor skeleton.
At first sight, this may appear to be a major blow to the plasticity-first hypothesis. And in some ways it is. However, rather than showing that morphological plasticity has been rather inconsequential in this adaptive radiation, Nathalie’s discussion of her results helps us sharpen our theories about the relationship between plasticity and evolution – something that we and others have argued is necessary. Step by step, this brings us one step closer to actually understanding how development shapes evolution.
Does evolution go where plasticity leads?
Mary-Jane West-Eberhard famously suggested that plasticity ‘takes the lead’ in adaptive evolution. But how can you tell if you are not there to see it happen? In a new paper in Evolution Letters, we show one way to tackle the problem – here is a quick summary of what we did and what we found.
We figured that we can look for signatures of plasticity by comparing the magnitude and direction of plastic responses to the environment with how locally adapted populations differ.
The literature on plant local adaptation is full of reciprocal transplant experiments that record both plasticity and adaptive divergence. We therefore used such data to test a number of hypotheses – and here is what we found:
First, the ancestral plastic response was generally well aligned with the difference between locally adapted phenotypes, making plasticity appear to ‘take the lead’ in adaptive evolution. Or – if you prefer – the plastic response of ancestors left a signature in the locally adapted phenotypes of descendants.
Second, plastic responses were sometimes more extreme than the locally adapted phenotypes. This can give the false impression that plasticity and evolutionary adaptation oppose each other: in fact, truly maladaptive plasticity was rare.
Third, although the signature of plasticity generally persists during local adaptation, genetic evolution modified trait combinations independently of plasticity.
We learnt a lot during this study. Perhaps the most important message is that we are a long way from understanding the relationship between plasticity and evolution – largely because very few studies are actually designed to test if evolution goes where plasticity leads. Hopefully this paper shows not just that this is a hypothesis that can be tested, but also help to clarify what kind of data that we need.
The first case of vertical transmission of a nematode from lizard mums to their offspring
The common perception of what scientist generally do probably includes making discoveries. But, in fact, the real life of a scientist mostly consists of collecting data, analyzing and interpreting data, reading and writing articles. Rarely are these activities interrupted by making discoveries, but it does happen.
While at Oxford, we began a project that investigated how lizard embryos in non-native populations in England cope with the cool climate. Unlike birds, who care for their offspring, most lizards bury their eggs in the soil. To study the adaptations that allow wall lizard embryos to develop at low temperatures, Nathalie spent most of the spring 2015 dissecting lizard eggs. Under the microscope, she opened the leathery egg shell with fine dissection tools and carefully detached the tiny embryo from the yolk. And she did this for hundreds of eggs.
The routine was disturbed one day by an odd discovery: in one of the embryos, Nathalie saw that something worm-like was wriggling in the brain of a baby-lizard! The discovery raised lots of excitement among her colleagues, but it also raised lots of questions: Why is the worm there and what is it actually? How did it get there – was it an accident or is this how the worm lives? Can the baby-lizard survive with worms wriggling in its brain?
Since we are lizard biologists rather than worm biologists, we teamed up with people who know more about nematodes with the aim to answer at least some of these questions. The result of this research can be read in our recent publication in the American Naturalist and we also wrote a little blog post that explains what answers we found. It is super cool and hopefully someone wants to pick this up and continue from where we left it!
Transposable elements unleashed
Mobile DNA sequences – transposable elements or TEs for short – are found in the genome of virtually all organisms. As their name implies, TEs can cut or copy themselves from one location in the genome to another. This can wreak havoc as insertion of TEs may interfere with gene regulation or in fact knock out entire genes. Cells therefore have mechanisms that prevent TEs from jumping, including DNA methylation and other epigenetic tools. Thus, TEs are not roaming freely through the genome, but are restricted from entering functionally important parts. Preventing TE invasion is particularly important when genes are regulated through spatial proximity to each other. The textbook example of this situation are the Hox genes, which are the key players in embryonic development with an ingenious mode of action: Hox genes are arranged in tight clusters and their position in the cluster defines their time and space of expression, and thus their effect on the patterning of the early embryo. It is therefore fitting that Hox gene clusters of mammals and other well-studied vertebrates have been found to be almost completely free of TEs. Nathalie’s new study reveals that Anolis lizards have broken this paradigm. Moreover, the invasion of TEs into Hox clusters of Anolis lizards can be linked to aberrant gene expression and increased rates of speciation.
Ever since the discovery of TEs, people have speculated about their evolutionary implications. One possible consequence of high TE activity is structural genomic variation. This may accelerate genomic incompatibility between populations; effectively making TEs engines of speciation.
Occasionally, TE insertions may also generate phenotypic novelty. As noted above, some genes are regulated through its proximity to other genes, which means that invasion of TEs can change expression of a number of genes simultaneously. Furthermore, since jumping TEs often drag along neighbouring genomic regions, they can translocate regulatory sequences that cause genes to be expressed in new cell types or at different stages in development.
While these are good reasons to expect TEs to promote evolution, examples are few and their role often appears idiosyncratic. An excellent group for a more systematic survey of TE-driven diversification are squamate reptiles, a group that includes lizards and snakes. Squamate genomes do not only appear particularly rich and variable in TEs, but their body plan is also highly malleable. Illustrative examples include the adaptive radiation of Anolis lizards and the repeated evolution of limbless and elongated bodies.
Nathalie decided to study how TEs have shaped the genomes, and in particular, the Hox clusters, of squamates. Her first surprise was to discover that lizards possess more Hox genes than all other tetrapods since they retained some genes that other lineages have ditched. The second surprise came when she looked at the TE content of Hox clusters. Despite the high TE content in their genomes, squamates follow other vertebrates in generally protecting their Hox clusters from TEs. But there was one exception: Nathalie found massive invasion of TEs in the Hox clusters of two out of three Anolis species, with TE contents almost as high as the average place in the genome.
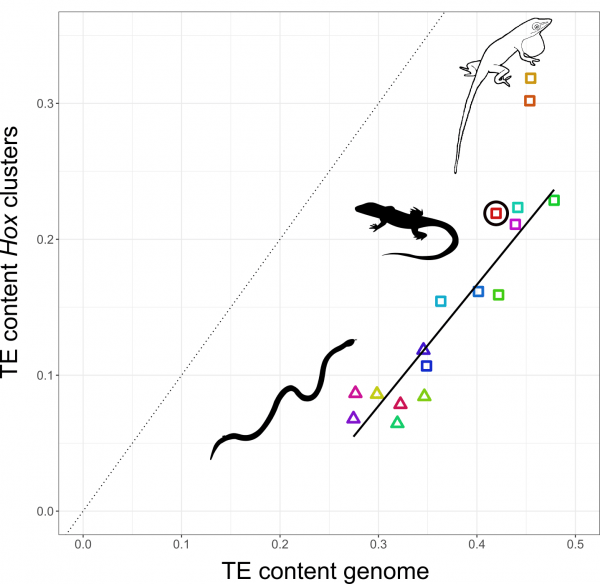
The relationship between TE content in Hox clusters relative to genome-wide TE content in squamate reptiles. All lizards and snakes restrict TEs from their Hox clusters down to roughly half the genome-wide average. However, two Anolis species show Hox clusters that are invaded by TEs, while a third Anolis species (black circle) follows the general trend.
Why in Anolis? Anolis lizards are famous in evolutionary biology due to their adaptive morphological radiation involving high rates of speciation – amassing close to 400 species. In a previous study Nathalie showed that Anolis lineages with more speciation events in the past have more TEs in their Hox clusters. This new genome-wide study reveals that this signature of speciation is indeed pronounced in Hox clusters: only the two Anolis species from amply speciating lineages exhibit unusually TE-rich Hox clusters, while a third species (Anolis auratus, black circle in figure above) follows the norm and keeps its Hox clusters relatively free from TEs. Looking in detail at genome-wide TE landscapes of these three Anolis species, Nathalie discovered that the two species with TE-rich Hox clusters had a larger population of young, more active TEs in their genomes. In addition, the inferred timing of peak activity of these TEs broadly coincided with past speciation events.
These results suggest that – during speciation events – TEs are unusually active and proliferate throughout the genome. As a result, even crucial regions such as Hox clusters become invaded. Subsequently, TEs are removed from Hox clusters by selection until a ‘healthy equilibrium’ of TE content relative to the genome-wide TE content is reached. This equilibrium appears highly conserved as the Hox clusters of almost all lizards and snakes contain close to 50% of the global TE content. This proposed model generates a number of predictions that can be tested with genomic data from lineages with variable rates of speciation.
How then do some Anolis species cope with having their Hox clusters invaded by TEs? Clearly, the inflation of Hox clusters – increasing the distance between genes – does not disrupt the patterning of the early embryo. Genes located at one end of the cluster remain expressed early in the head of the embryo, while genes located at the other end are expressed late in the tail. However, the successive activation of Hox genes predicts that disruption, if occurring at all, should be most pronounced towards the end of the Hox clusters. Nathalie found that this indeed is the case: one out of four Hox13 genes showed aberrant expression in the two Anolis species with TE invaded Hox clusters, but this gene was expressed as ‘normal’ in other Anolis and more distantly related lizards.
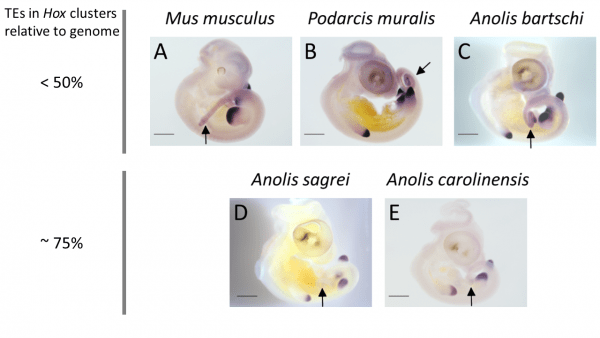
Expression patterns of the posterior Hox gene HoxD13 are showing variation between species with low and high TE content in their Hox cluster: while expression in limb buds is conserved, expression in tail tissue (black arrows) is missing in species with high TE content in their Hox clusters (A. sagrei and A. carolinensis).
Nathalie’s study reveals that, despite being THE textbook example of our conserved developmental toolkit, Hox genes can be tinkered with. What is more, the TE invasion of Hox clusters appear to be intimately linked to diversification. Now that Anolis lizards have shown us that it can happen, perhaps they can also show us why it happens and how.
The original version of this blog post was published on the Evolution Letters Editors’ Blog https://evolutionletters.wordpress.com/2019/08/06/the-role-of-mobile-genetic-elements-in-evolution-and-development/
Hot off the press!
 Our book Evolutionary Causation has arrived! It not only looks great, but it also features some really original and exciting ideas by fantastic people. Thanks to all the authors and to the KLI and the MIT Press for making it happen.
Our book Evolutionary Causation has arrived! It not only looks great, but it also features some really original and exciting ideas by fantastic people. Thanks to all the authors and to the KLI and the MIT Press for making it happen.
Interested in the content? Here is a list of titles and authors:
1 Evolutionary Causation
Tobias Uller and Kevin N. Laland
2 Causality and the Role of Philosophy of Science
Massimo Pigliucci
3 Understanding Bias in the Introduction of Variation as an Evolutionary Cause
Arlin Stoltzfus
4 The Shape of Things to Come: Evo Devo Perspectives on Causes and Consequences in Evolution
Armin P. Moczek
5 Incorporating the Environmentally Sensitive Phenotype into Evolutionary Thinking
David I. Dayan, Melissa A. Graham, John A. Baker, and Susan A. Foster
6 Genotype-Environment Interaction and the Unscripted Reaction Norm
Sonia E. Sultan
7 Understanding Niche Construction as an Evolutionary Process
Kevin N. Laland, John Odling-Smee, and Marcus W. Feldman
8 Biological Dynamics and Evolutionary Causation
Renée A. Duckworth
9 The Causes of a Major Transition: How Social Insects Traverse Darwinian Space
Heikki Helanterä and Tobias Uller
10 Are Developmental Plasticity, Niche Construction, and Extended Inheritance Necessary for Evolution by Natural Selection? The Role of Active Phenotypes in the Minimal Criteria for Darwinian Individuality
Richard A. Watson and Christoph Thies
11 The Paradox of Population Thinking: First-Order Causes and Higher-Order Effects
Denis M. Walsh
12 Ontology, Causality, and Methodology of Evolutionary Research Programs
Jun Otsuka
13 A Darwinian Dream: On Time, Levels, and Processes in Evolution
Arnaud Pocheville
14 Decoupling, Commingling, and the Evolutionary Significance of Experiential Niche Construction
Lynn Chiu
15 Biological Information in Developmental and Evolutionary Systems
Karola Stotz