Impressions from field work 2022

DNA methylation is inherited in water fleas
As the EU-funded project IDEAL came to a close, Tobias and Bas Heijmans looked for a good scientific reason to stay in touch. Tobias’ plans to work on water fleas seemed like a promising way to collaborate. The only problem was that the project was not funded, but a Wallenberg Academy Fellowship soon made it possible to get going.
Thanks to Reinder Radersma, Elmar Tobi and Amanda Raine we got the methods working, designed an experiment and collected data. Our aim was modest: we simply wanted to test if – and to what extent – environmentally induced DNA methylation really was heritable in Daphnia. If it was, lots of interesting things could be done. If it wasn’t, we would not need to worry about missed opportunities to learn something new. Either way, we’d all learn new stuff and have fun.
As it happens, various things slowed us down. Reinder and PhD student Alex pushed on with other experiments, which suggested that – at least as far as adaptation to toxic cyanobacteria goes – the importance of non-genetic inheritance may have been exaggerated. All of us were busy with other projects and, to be honest, the lizards were just so much fun. The DNA methylation project entered a dauer stage – inactive, motionless, yet alive.
It could have stayed that way were it not for a grant from SciLifeLab (thanks!), and Nathalie took up the challenge as Reinder began his new job. Together with Louella, Markus and Björn, Nathalie demonstrated that our ideas were not so off the mark after all – environmentally induced DNA methylation really seemed to be inherited in water fleas.

Overlap between differentially methylated positions (DMPs) between the three generations in all four stressors.
Exposing mothers to one of three natural stressors – high temperature, zinc or cyanobacteria – induced a similar number of differentially methylated CpGs (DMPs) in their offspring. These DMPs were not randomly distributed across the genome, but almost exclusively found in gene bodies. Previous work suggests that this is how DNA methylation affects gene expression in invertebrates.
It turned out that about half of the DMPs persisted into the F4 generation. We were surprised at this high proportion, but the results were robust to our attempts to explain them away.
Since we had included a treatment with a de-methylating drug, we could also show that Daphnia in fact has a reliably machinery to re-methylate the genome. Thus, the environmentally induced DMPs to a very large extent escapes the ‘normal’ re-methylation.

5-Azacytidine strongly de-methylates CpG sites in the F1 generation, but methylation marks are re-set already in generation F2.
It is tempting to interpret these results as evidence that inheritance of DMPs has a biological function rather than being a system failure. Whether or not that is the case, the environmentally induced DMPs can certainly have biological consequences since they were enriched in genes with particular functions – functions that were specific to the stressors.
Looking back, it may seem discouraging that it took ten years from idea to paper for a ‘simple’ proof-of-principle study. Well, sometimes it is just like that – ideas are cheap, putting them into practice is what’s hard. But now we know that it is worthwhile to study the ecological and evolutionary consequences of epigenetic inheritance in water fleas. New molecular biology methods and field experimental set-ups make this a pretty exciting project.
You can read the paper published in iScience here.
Dorsal pattern formation in Anolis lizards
Readers of this blog will be well aware of the conspicuous variation in dorsal color patterns of Anolis lizards – think of the spots of A. sabanus, or the bands of A. transversalis (both worthy a google search if you happen to be unfamiliar with them). Such patterns are particularly interesting when several patterns exist within a single population – a polymorphism. This is the case in A. sagrei where females have either a chevron-like pattern, same as all males of this species, or a diamond-like pattern on their backs.
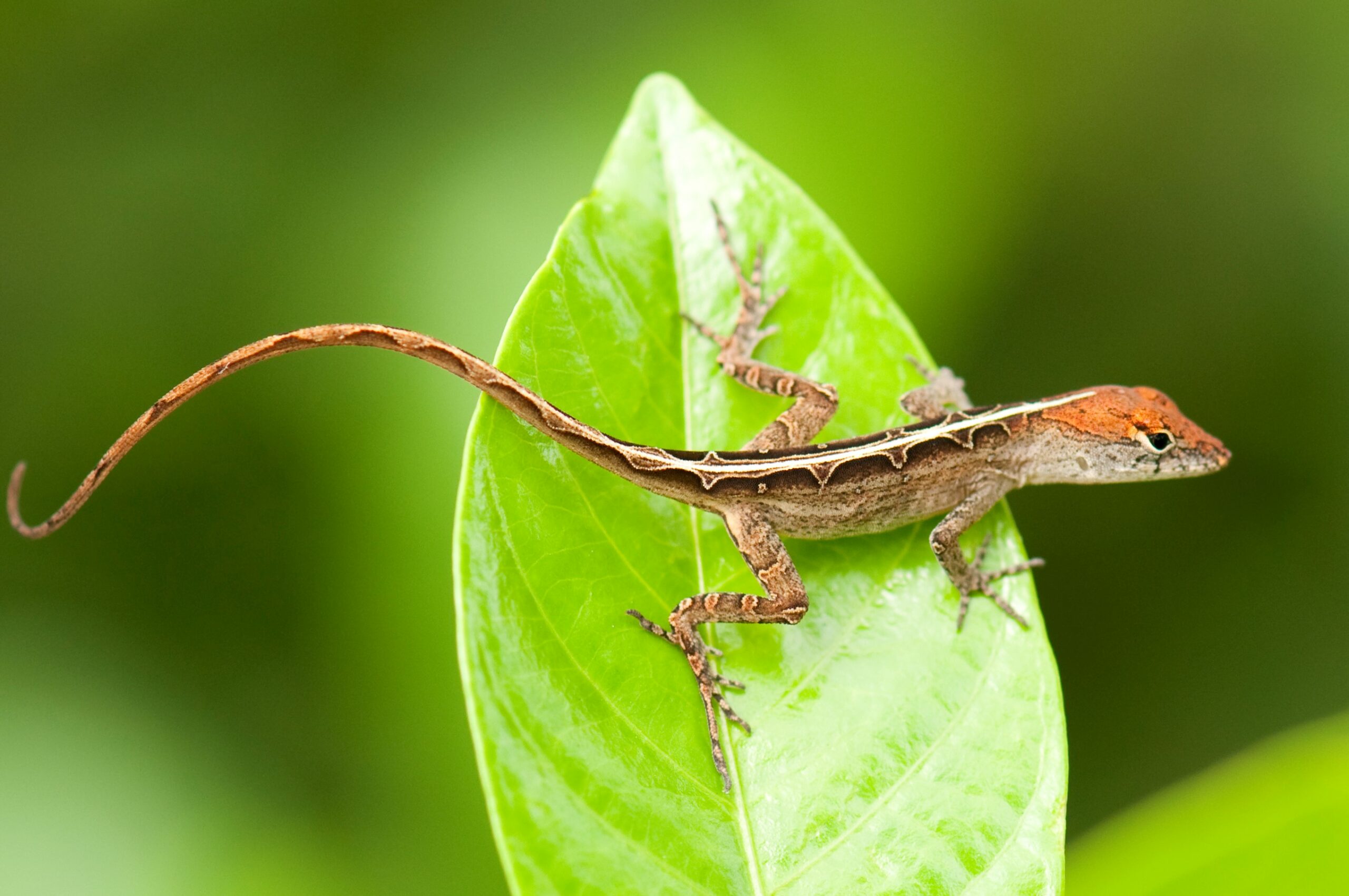

Picture of a female A. sagrei with the diamond pattern.
In 2016, I visited some A. sagrei populations in Florida and became interested in this female-limited color pattern polymorphism. Being a developmental biologist, what puzzled me most was not if selection can maintain multiple patterns, but how the two patterns actually develop.
On the one hand, the difference between the two patterns ought to have a simple genetic basis. Diamond- and chevron-females mix within the same population and the two morphs are discrete (although there is a continuous variation within both morphs, and the diamonds sometimes look more like a band [1]). On the other hand, the developmental biology of pattern formation is far from trivial. Here is a little teaser of the complexity: in vertebrates, pigment cells are derived from neural crest cells that originate along the dorsal midline of the early embryo. Once detached from the crest of the neural tube, pigment cell precursors migrate towards their final destination in the epidermis, where they differentiate into pigment cells – xanthophores, melanophores and iridophores. What these cells do and how they organize themselves are what gives rise to patterns. And this does not seem to be very simple at all. So how can we reconcile a simple genetic basis with the complex developmental biology of pattern formation of diamonds and chevrons?
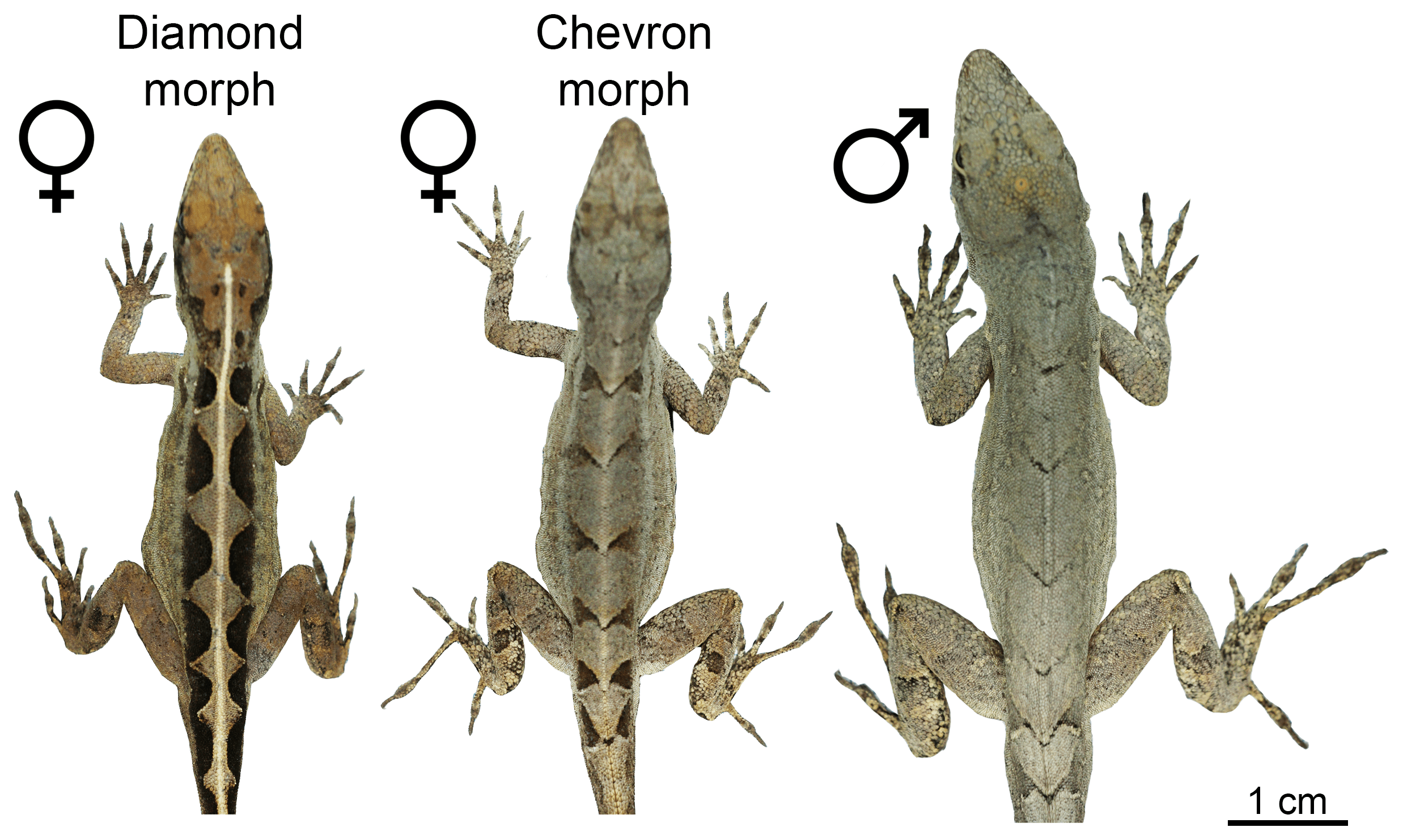

The two female morphs and a male of A. sagrei.
Simply out of curiosity, we set out to solve this puzzle. Thanks to our anole breeding group at Lund University, we already had three generations of A. sagrei of both morphs that we could use to address the pattern of inheritance of diamonds and chevrons, and to identify the underlying gene(s). Our intuition about the simple genetic architecture of the polymorphism held true, and we identified a single Mendelian locus that perfectly segregated with the morph patterns.
Surprisingly though, the locus was not on a sex chromosome, so this could not explain why only females are polymorphic. Instead, the identity of one of the two genes located at the Mendelian locus pointed to a solution: the estrogen receptor 1 gene is crucial for the development of female traits and therefore expressed at higher levels in females compared to males. Because of the close physical proximity of the two genes at the Mendelian locus, also the second gene shows a female-biased expression pattern. So while the estrogen receptor 1 gene does not explain the pattern differences, it explains why the polymorphism is only present in females.
But what is this second gene at the Mendelian locus, and how does it explain the difference in patterning? It is a gene encoding the coiled-coil domain-containing protein 170, or CCDC170. Not exactly a usual suspect of color pattern formation. In fact, not much is known about this gene, but the cancer literature had demonstrated that it codes for a structural protein that regulates the migratory capacity of cells (mutations in CCDC170 can make cancer cells migratory and are associated with an aggressive type of breast cancer). The proteins encoded by the diamond and chevron alleles were predicted to form CCDC170 proteins with structural differences. This is likely to affect the function of the protein – perhaps by influencing migratory behaviors of pigment cell precursors.
Testing this hypothesis is easier said than done. As a first proof-of-principle, we used an in silico modelling approach to test if tinkering with the migratory capacity of cells can switch a system from generating chevron-like to diamond-like patterns. Miguel Brun-Usan is a magician with cell-based computer modelling and his trick is to create computer models that are complex enough to capture real biological phenomena, yet simple enough to allow us to trace and understand what is going on. To get to the bottom of the diamond/chevron morph differences, he implemented cells on a growing epithelium with a Turing-type mechanism consisting of a gene regulatory network (GRN) with up to five genes. By running a number of experiments (yes, computer scientists also run experiments, I learned), Miguel could demonstrate that some GRNs generated back patterns that very much resembled those of real lizards. Moreover, he found that modifying a single gene in the GRN that regulates the migratory capacity of cells is sufficient to switch the system from generating chevrons (if migration is switched ON) to diamonds (if migration is switched OFF).
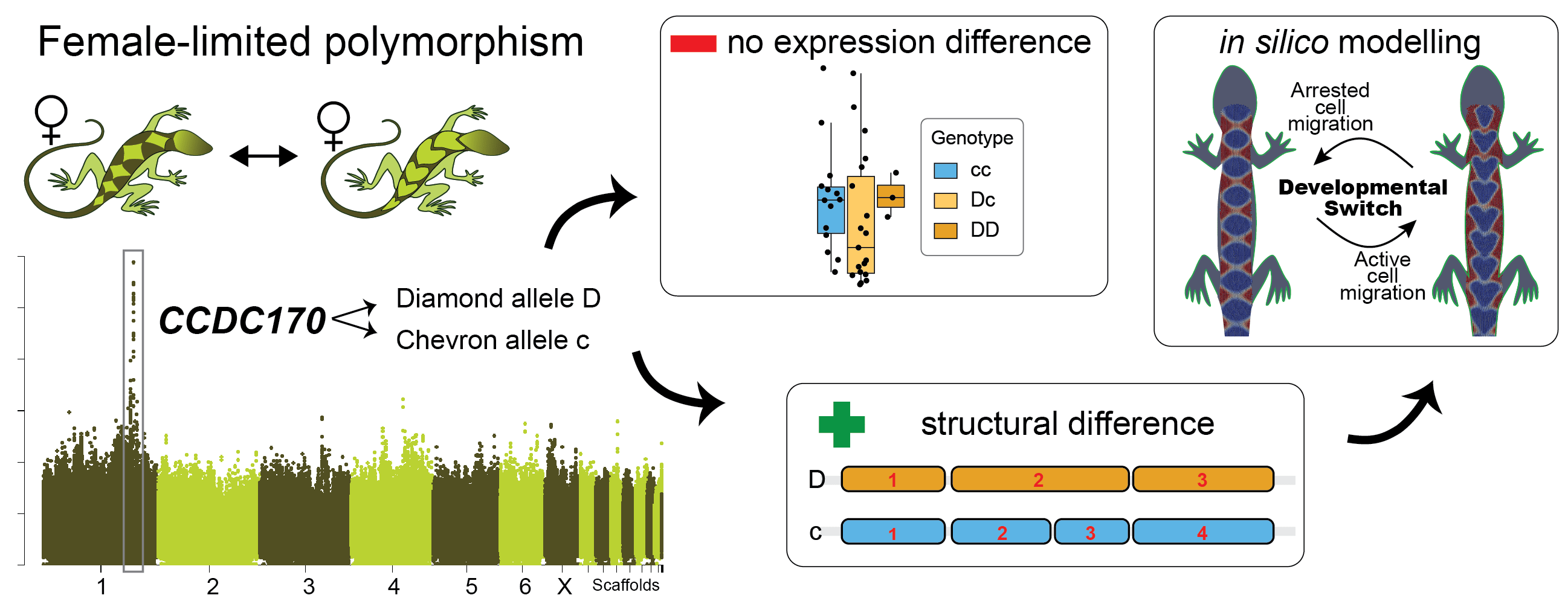
A brief graphical abstract of our study.
So it seems like what reconciles the simple genetic basis of the polymorphism with the complex process of pattern formation is that the underlying developmental system exhibits a high degree of controllability. Simply modifying migratory capacities of cells by tinkering with one gene in the core GRN leads to changes in collective cell migration, visible as a distinct and new dorsal color pattern. Such high controllability could explain the evolvability of dorsal color patterns and result in high turn-over rates between patterns, as beautifully demonstrated by geckos [2]. I am excited to continue this research to see if the model can be substantiated mechanistically. I would also like to test if convergent evolution of diamond patterning in different Anolis species is underpinned by convergent developmental and genetic mechanisms. If you are interested in joining our research group, please get in touch (nathalie.feiner@biol.lu.se).
You can read the full article here:
Feiner, N., M. Brun-Usan, P. Andrade, R. Pranter, S. Park, D. B. Menke, A. J. Geneva, and T. Uller. 2022.
A single locus regulates a female-limited color pattern polymorphism in a reptile.
Science Advances 8:10 DOI: 10.1126/sciadv.abm2387
Other references:
1. Moon, R.M., and Kamath, A. (2019). Re-examining escape behaviour and habitat use as correlates of dorsal pattern variation in female brown anole lizards, Anolis sagrei (Squamata: Dactyloidae). Biol J Linn Soc 126, 783-795.
2. Allen, W.L., Moreno, N., Gamble, T., and Chiari, Y. (2020). Ecological, behavioral, and phylogenetic influences on the evolution of dorsal color pattern in geckos. Evolution 74, 1033-1047.
The macroevolutionary consequences of island hopping
Natural historians have known for a long time that islands often harbour an extraordinary amount of biodiversity. One reason is that lineages that colonize islands can exploit open ecological niches and therefore diversify along new evolutionary trajectories. So did Anolis lizards following their arrival to the Caribbean. A range of habitat specialists has evolved repeatedly on different islands, creating a structured diversity of morphologies. Since one Anolis clade has remained on the mainland, and one clade has returned the mainland, this group of lizards presents a kind of before/after ‘island evolution’ experimental setting that can help us understand how evolution works.
Our new study, published in Nature Communications, took advantage of this setting to find out if morphological diversification proceeded in a different way following colonization of the Caribbean islands compared to the lizards that stayed on the mainland. This would be predicted if there were lots of ecological opportunities on islands. Furthermore, we wanted to know if the clade that returned to the mainland evolved along similar trajectories as their distant mainland relatives, or if they followed more closely their Caribbean relatives. Finally, we also wanted to test if evolution on the Caribbean islands was faster, more punctuated, and led to higher morphological diversity than on the mainland, again expected if lizards encountered a lot of empty niches on islands. To answer these questions, Nathalie and Illiam travelled to the Nanoscale Research Facility at the University of Florida to collect CT-scans of more than 700 individuals of 271 Anolis species. Back home they quantified the morphology of the parts of the skeleton that determine how the lizards navigate their habitat – limb bones, hip and shoulder girdles. Interestingly, we found that the evolutionary modularity – the covariation of parts of the locomotor skeleton across the evolutionary tree – was different between the clades: while the lizards whose ancestors had never set foot on an island showed a tight covariation between the fore- and hindlimbs, Anolis evolution on islands followed a trajectory with much tighter covariation between limbs and their respective girdles (forming a front and a hind module). What is more, the evolutionary modularity that was seen across island lizards was also seen in the mainland re-colonizers.
What does this mean? The answers are hidden in the past, of course, but we speculate that the exploitation of ecological opportunities following colonization of the islands led to a profound change in how limbs and girdles develop and grow together during ontogeny. That is, adaptation on islands may have changed how limbs and girdles vary together, and this change persisted even after lizards recolonized the mainland. This meant that lizards in the clade that recolonized the mainland produced very similar morphologies to those seen on the islands, but failed to explore some parts of the morphospace occupied by their other mainland relatives. If we compare the island lizards with the clade that returned to the mainland, we find that a general prediction from island biogeography holds true – island lizards show faster and more variable evolutionary rates and greater morphological variation. That is, while the patterns of diversification of the locomotor skeleton is similar, the mainland lizards produce only a subset of the morphologies seen on the islands. The same is not true, however, if we compare the morphology of island lizards to the morphology of lizards in the ‘primary’ mainland clade. This mainland clade is equally diverse as the lizards on islands, and it contains morphologies that are not seen on the islands. Perhaps this means that comparing diversification patterns of island and mainland groups is only ‘fair’ if the two groups share the same variability. Or perhaps it reflects historical differences in the ecological opportunities for adaptive diversification in the two mainland clades.
We do not yet have all the answers, and many will forever remain unknown. However, we do suggest that these macroevolutionary trends of the locomotor skeleton in Anolis illustrate that adaptation to ecological opportunities on islands can have profound effects on trait development, with lasting effects on patterns of morphological diversification. To put this hypothesis to the test, we need to study how the parts of the locomotor skeleton vary together within species, and do this for several species across the different clades. A big task, but not impossible.
New paper throws doubt on a popular idea
The rock-paper-scissors story of the side-blotched lizards was beautiful. Many lizards have similar colour polymorphism, and quite a few PhD projects and postdoc years have been spent testing the idea in other species. One of the most obvious candidates are wall lizards, where several species come with clearly distinct white, yellow and orange/red throats and bellies. The early reports were – as they often are – encouraging. But researchers throughout Europe grew increasingly worried that the phenotypic differences between red, yellow and white animals at best were small and inconsistent or, at worst, entirely absent.
Alternative mating strategies tend to be manifested behaviourally. We therefore decided to release common wall lizards of the three main colour morphs in large outdoor enclosures in Moulis and observe what they actually do. The prediction is simple: if the morphs signal relevant aspects of sociosexual behaviour, such differences should be evident in how the lizards interact with one another, or who they chose to mate with.
We had done similar studies before, both in Moulis and in Oxford, and it works like a dream. As long as there are people truly dedicated to watching lizards, that is. Thankfully, the lead author of the study – Javier Abalos – is very patient and, quite frankly, obsessed with lizard watching.
With a huge amount of behavioural data collected, home ranges quantified, and paternity secured, Javier could conclusively say that there is nothing going on. Nothing, Nada, Inte ett dugg.
There is of course always the possibility that lizards do something else when they go about their business outside of field enclosures. So Javier and another obsessive lizard watcher – Guillem Perez I de Lanuza – added a long term field data set to the mix. Again, no evidence that males or females of different colour morphs behave differently, or that they modify their behaviour depending on the morph of other individuals.
Conclusion: Whatever these colours mean, they certainly do not signal differences in social or sexual behaviour. At least not in this lineage of wall lizards. You can read the full story here.
The problem, of course, is that it is very difficult to prove that there is nothing going on. It is also very unpopular when data fail to support a good old adaptive story. So reviewers try hard to come up with reasons for rejection – it can almost be endearing to read attempts to rescue a favourite hypothesis. But efforts like those from Allen Moore and the editors of Ecology & Evolution ensure a swift publication of unpopular findings. Chapeau to the editor for their frank evaluation of the reviewer comments from a different journal. Such encouragement is especially important for early career researchers so they won’t be put off by people who care more about their ego than about what the results say.
Nevertheless, at the end of the day, there must be some kind of explanation for why these morphs persist. They are even found in several other wall lizard species. But after Javier’s results you would have to be an optimist beyond belief to think they signal alternative behavioural strategies. If you have different ideas and want to test them, please don’t hesitate to get in touch.



