Readers of this blog will be well aware of the conspicuous variation in dorsal color patterns of Anolis lizards – think of the spots of A. sabanus, or the bands of A. transversalis (both worthy a google search if you happen to be unfamiliar with them). Such patterns are particularly interesting when several patterns exist within a single population – a polymorphism. This is the case in A. sagrei where females have either a chevron-like pattern, same as all males of this species, or a diamond-like pattern on their backs.

Picture of a female A. sagrei with the diamond pattern.
In 2016, I visited some A. sagrei populations in Florida and became interested in this female-limited color pattern polymorphism. Being a developmental biologist, what puzzled me most was not if selection can maintain multiple patterns, but how the two patterns actually develop.
On the one hand, the difference between the two patterns ought to have a simple genetic basis. Diamond- and chevron-females mix within the same population and the two morphs are discrete (although there is a continuous variation within both morphs, and the diamonds sometimes look more like a band [1]). On the other hand, the developmental biology of pattern formation is far from trivial. Here is a little teaser of the complexity: in vertebrates, pigment cells are derived from neural crest cells that originate along the dorsal midline of the early embryo. Once detached from the crest of the neural tube, pigment cell precursors migrate towards their final destination in the epidermis, where they differentiate into pigment cells – xanthophores, melanophores and iridophores. What these cells do and how they organize themselves are what gives rise to patterns. And this does not seem to be very simple at all. So how can we reconcile a simple genetic basis with the complex developmental biology of pattern formation of diamonds and chevrons?
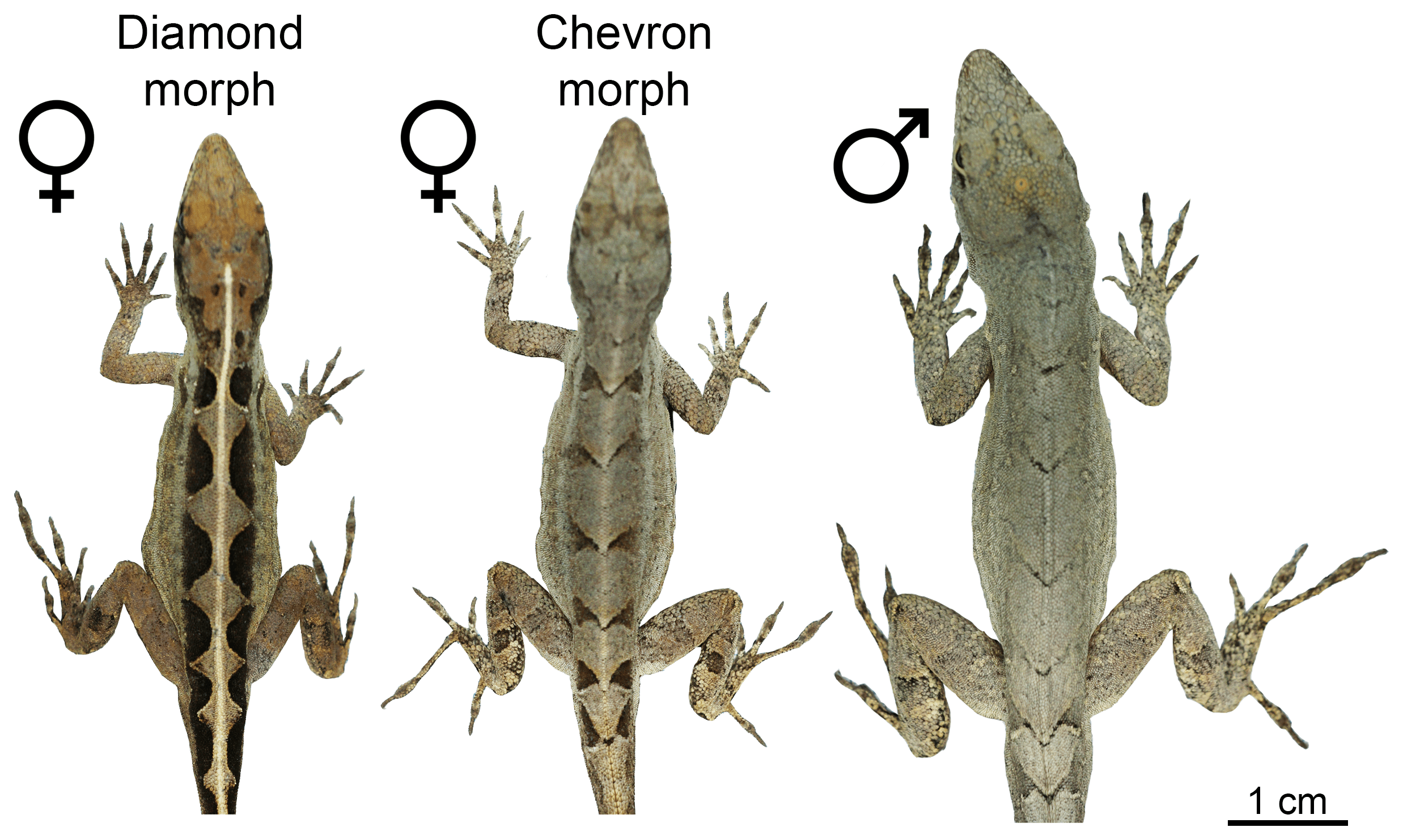

The two female morphs and a male of A. sagrei.
Simply out of curiosity, we set out to solve this puzzle. Thanks to our anole breeding group at Lund University, we already had three generations of A. sagrei of both morphs that we could use to address the pattern of inheritance of diamonds and chevrons, and to identify the underlying gene(s). Our intuition about the simple genetic architecture of the polymorphism held true, and we identified a single Mendelian locus that perfectly segregated with the morph patterns.
Surprisingly though, the locus was not on a sex chromosome, so this could not explain why only females are polymorphic. Instead, the identity of one of the two genes located at the Mendelian locus pointed to a solution: the estrogen receptor 1 gene is crucial for the development of female traits and therefore expressed at higher levels in females compared to males. Because of the close physical proximity of the two genes at the Mendelian locus, also the second gene shows a female-biased expression pattern. So while the estrogen receptor 1 gene does not explain the pattern differences, it explains why the polymorphism is only present in females.
But what is this second gene at the Mendelian locus, and how does it explain the difference in patterning? It is a gene encoding the coiled-coil domain-containing protein 170, or CCDC170. Not exactly a usual suspect of color pattern formation. In fact, not much is known about this gene, but the cancer literature had demonstrated that it codes for a structural protein that regulates the migratory capacity of cells (mutations in CCDC170 can make cancer cells migratory and are associated with an aggressive type of breast cancer). The proteins encoded by the diamond and chevron alleles were predicted to form CCDC170 proteins with structural differences. This is likely to affect the function of the protein – perhaps by influencing migratory behaviors of pigment cell precursors.
Testing this hypothesis is easier said than done. As a first proof-of-principle, we used an in silico modelling approach to test if tinkering with the migratory capacity of cells can switch a system from generating chevron-like to diamond-like patterns. Miguel Brun-Usan is a magician with cell-based computer modelling and his trick is to create computer models that are complex enough to capture real biological phenomena, yet simple enough to allow us to trace and understand what is going on. To get to the bottom of the diamond/chevron morph differences, he implemented cells on a growing epithelium with a Turing-type mechanism consisting of a gene regulatory network (GRN) with up to five genes. By running a number of experiments (yes, computer scientists also run experiments, I learned), Miguel could demonstrate that some GRNs generated back patterns that very much resembled those of real lizards. Moreover, he found that modifying a single gene in the GRN that regulates the migratory capacity of cells is sufficient to switch the system from generating chevrons (if migration is switched ON) to diamonds (if migration is switched OFF).
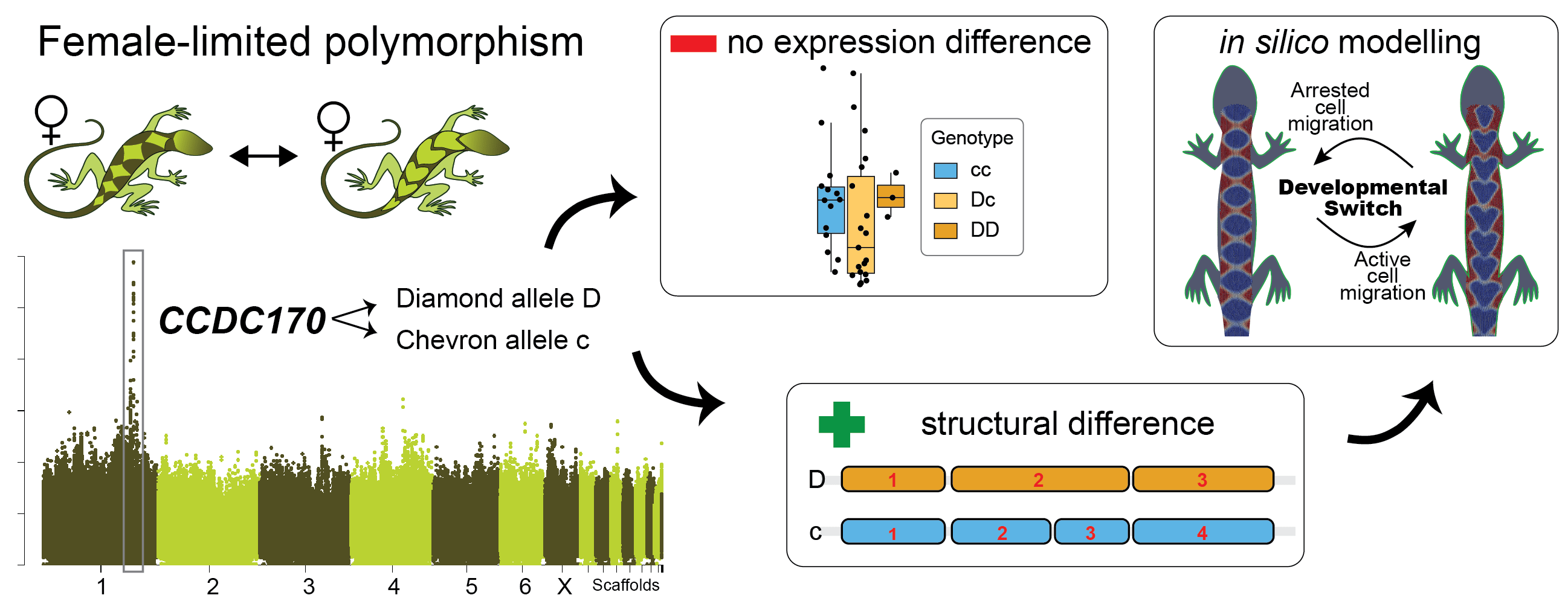
A brief graphical abstract of our study.
So it seems like what reconciles the simple genetic basis of the polymorphism with the complex process of pattern formation is that the underlying developmental system exhibits a high degree of controllability. Simply modifying migratory capacities of cells by tinkering with one gene in the core GRN leads to changes in collective cell migration, visible as a distinct and new dorsal color pattern. Such high controllability could explain the evolvability of dorsal color patterns and result in high turn-over rates between patterns, as beautifully demonstrated by geckos [2]. I am excited to continue this research to see if the model can be substantiated mechanistically. I would also like to test if convergent evolution of diamond patterning in different Anolis species is underpinned by convergent developmental and genetic mechanisms. If you are interested in joining our research group, please get in touch (nathalie.feiner@biol.lu.se).
You can read the full article here:
Feiner, N., M. Brun-Usan, P. Andrade, R. Pranter, S. Park, D. B. Menke, A. J. Geneva, and T. Uller. 2022.
A single locus regulates a female-limited color pattern polymorphism in a reptile.
Science Advances 8:10 DOI: 10.1126/sciadv.abm2387
Other references:
1. Moon, R.M., and Kamath, A. (2019). Re-examining escape behaviour and habitat use as correlates of dorsal pattern variation in female brown anole lizards, Anolis sagrei (Squamata: Dactyloidae). Biol J Linn Soc 126, 783-795.
2. Allen, W.L., Moreno, N., Gamble, T., and Chiari, Y. (2020). Ecological, behavioral, and phylogenetic influences on the evolution of dorsal color pattern in geckos. Evolution 74, 1033-1047.